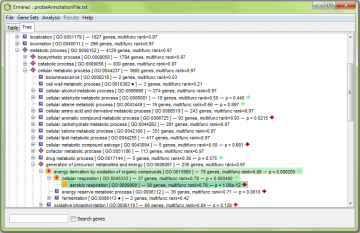
- Double-click on a node to view the details. Single-click on a node to expand its immediate children.
- Right-click (MacOS:option-click) on a node to get more tools.
- If you have more than one result set loaded, you can choose the one to show in the tree using the “Results” menu.
- Terms which have no genes in your annotations are hidden by default but can be shown using the pop-up menu.
- Your user-defined groups are shown as a separate “pseudo-branch” of the tree.
What the icons mean
The icons indicate the state of a node, as well as that of its children. This makes it easier to see that even if a term is not “significant”, it has a child term that does. To expose the sub-tree at a node, right-click and select “Expand this node”.
 – The group is not significant, and has no significant children
– The group is not significant, and has no significant children – The group is not significant, but it has significant children
– The group is not significant, but it has significant children – The group is significant, but has no significant children
– The group is significant, but has no significant children – The group is significant, and it has significant children
– The group is significant, and it has significant children
Where “significant” means after multiple test correction.
Limitations
- Only one path to the root is shown per GO term. Other immediate parents of each term are only shown in the tool-tip for the term. To see an accurate representation of the GO hierarchy, use the GO consortium’s web site, also accessible through the context (pop-up) menu.
- Attempting to expand large sections of the tree is slow; you may see an error message saying it was cancelled.